Engineering Success
Iterative optimization of a scalable biomanufacturing process for photothermal therapy-optimized nanoparticle production
CYCLE №1
Identify the optimal ratio of gold:silver salts for synthesis of photothermal therapy (PTT)-optimized nanoparticles
CYCLE №2
Identify the growth medium for obtaining the optimal bioreducing mix for the production of PTT-optimized nanoparticles
CYCLE №3
Test the influence of Escherichia coli genes copA, napA, cueO, and melA on nanoparticle synthesis
CYCLE №4
Test if growth on a nitrate-containing medium benefits the bioreducing capacity of the supernatant used for nanoparticle production
Overview
With Binanox, we set ourselves the goal of establishing a novel, scalable process for the production of bimetallic nanoparticles using a cell-free system with metal-reducing proteins. Within the scope of our project, we approach process design from a scale-up perspective by focusing on understanding and optimizing the process at a lab scale first, then proceeding to pilot and factory levels. Therefore, we employed the Design-Build-Test-Learn engineering cycle framework to ensure that our system is optimized at every iteration. On this page, you will find a description of how four such cycles guided the creation of Binanox.


Introduction
To approach process design efficiently, we started by looking at the monumental efforts of our predecessors1. Due to similarities in reaction conditions, we took green chemical production methods as a reference point for our process design. Namely, the synthesis of nanoparticles with ascorbic acid, as outlined by Cheng et al. 2 was deemed the most representative of our aspirations. The authors obtained star-shaped nanoparticles with a plasmon peakinfo at 800 nm. This fits almost all our prerequisites (read more about this on the Project Description page), as these properties are optimal for use in photothermal therapy (PTT). However, this methodology does not yield bimetallic nanoparticles. We aimed to make this possible with our biomanufacturing process by using this synthesis methodology as the starting point for our process design.
Ascorbic acid is among the standard methodologies for the production of nanoparticles, but it poses several difficulties in scaling the production industrially3. At the same time, the most common chemical reducing agent for industrial production is sodium borohydride, which produces toxic, pyrophoric and flammable compounds upon oxidation (B2H6 and H2 gasses)4. Using biological reducing agents, we want to provide a safe, scalable, and efficient methodology for producing bimetallic star-shaped nanoparticles5.
Cycle 1. Determining the optimal HAuCl4:AgNO3 ratio for PTT-optimized nanoparticles production with ascorbic acid

Motivation
Binanox is a biomanufacturing system with many variables, which impels preliminary simplification to learn more about the process.

Design
Investigate a more simple synthesis method before proceeding to a complex, bioreducing system.

Build & Test
Use Box-Behnken Design for modeling nanoparticle synthesis with ascorbic acid.

Learn
Optimal gold:silver salts ratio for synthesis of PTT-optimal nanoparticles in biologically-relevant conditions is 1.85:1.
Design
First, we decided to identify the optimal conditions for chemical nanoparticle synthesis using ascorbic acid. This way we could investigate several parameters in a more simple setting before moving on to a complex biological system. Additionally, to bridge the gap between chemical and biological synthesis, the reduction with ascorbic acid was conducted at pH 7 instead of pH 3, as done in the reference paper2. This was done under the presumption that optimal functionality of reducing enzymes present in Escherichia coli (E.coli) is achieved at neutral pH, as it corresponds to the pH of optimal growth in this bacterium6. As this methodology deviates from the original paper, the ascorbic acid experiments were repeated at biologically relevant pH to obtain an optimal gold:silver (HAuCl4:AgNO3) ratio for this pH range.
We used Box-Behnken Design to minimize the number of experiments necessary to identify the optimal HAuCl4:AgNO3 ratio for the synthesis of nanoparticles with a plasmon peak at 800 nm. For a more detailed description of Box-Behnken Design, you can refer to the Modeling page. We could find the optimum for three variables in just 15 experiments instead of 27. The three variables were (1) the concentration of AgNO3 (2) the concentration of HAuCl4, and (3) pH. The latter was varied between 5 and 9 to test the influence of pH on the formation of nanoparticles under biologically relevant pH conditions6.
Build & Test
After synthesis, the 450-1000 nm absorption spectrum was measured for each sample. These spectra were used as input for the model, which then returned a correlation between the three variables and the absorbance, displayed as an interactive slider in Fig. 3. This yielded a prediction of the spectrum, while the main optimization objective was a high absorbance at 800 nm. You can refer to the Modeling page for a detailed discussion of the methodology and the results of this experiment.
If you enjoy tweaking this slider, check out our Modeling page! Here, you can check the effect of pH, temperature, and gold concentration on the absorbance of nanoparticles in a cell-free system. This tool can be useful for setting up a production process according to prerequisites. This is an important step in the upscaling of the biomanufacturing process.
Learn
The model predicted an optimal molar ratio of HAuCl4:AgNO3 = 1.80:1 at a pH of 6.1. However, as will be elaborated in Cycle 2, Mueller Hinton (MH) Broth was shown to be an optimal growth medium for nanoparticle production. The pH of MH Broth is 7.3, thus the model was adjusted to maximize A800 nm for this pH value. This yielded a ratio of 1.85:1. Subsequently, the spiky morphology of the prepared nanoparticles was confirmed using TEM (Fig. 4). This “golden ratio” was used for all subsequent experiments with the biological system, because the produced nanoparticles in this experiment had the correct shape and size to observe plasmon resonance at 800 nm.

Cycle 2. Getting the right biological system for nanoparticle production

Motivation
A literature protocol for short nanoparticle synthesis with MH broth did not work 7.

Design
Adjust the nanoparticle synthesis protocol to span over 24h and test 4 common growth media.

Build & Test
Test nanoparticle synthesis with lysed and non-lysed supernatants of wildtype E. coli grown in 4 broths: LB, LB low salt, MH, TSBS.

Learn
Non-lysed MH broth yielded the best performance in PTT-optimized nanoparticle synthesis.
Design
The next step in developing the biomanufacturing system was establishing which growth medium allows the model organism E. coli BL21 to produce the most effective blend of reducing agents. Extensive literature reviews show that the reducing agents play a crucial role in the size and morphology of the produced nanoparticles 4, which demands fine-tuning the bioreducing system.
Gurunathan et al.8 has shown the capacity of E. coli supernatant to produce silver nanoparticles from AgNO3 within five minutes7. The supernatant was obtained after 24h of growth in MH broth. After unsuccessfully attempting this conveniently short synthesis method, another paper stating that nanoparticle synthesis occurred in 24h was consulted8. Preliminary tests showed that this timespan was indeed sufficient. Therefore, after this point, all biochemical synthesis was performed over 24h.
To assess the optimal media for bacterial growth, we performed nanoparticle synthesis with supernatants of four different media: LB, LB Low Salt, MH, TSBS. Another variable tested was whether it is better to lyse the cells used for nanoparticle production. Lysing of the cells can be achieved by sonicating the cell liquid culture before separating the supernatant from the pellet. The Results page further discusses the choice of media and sonication settings.
Build & Test
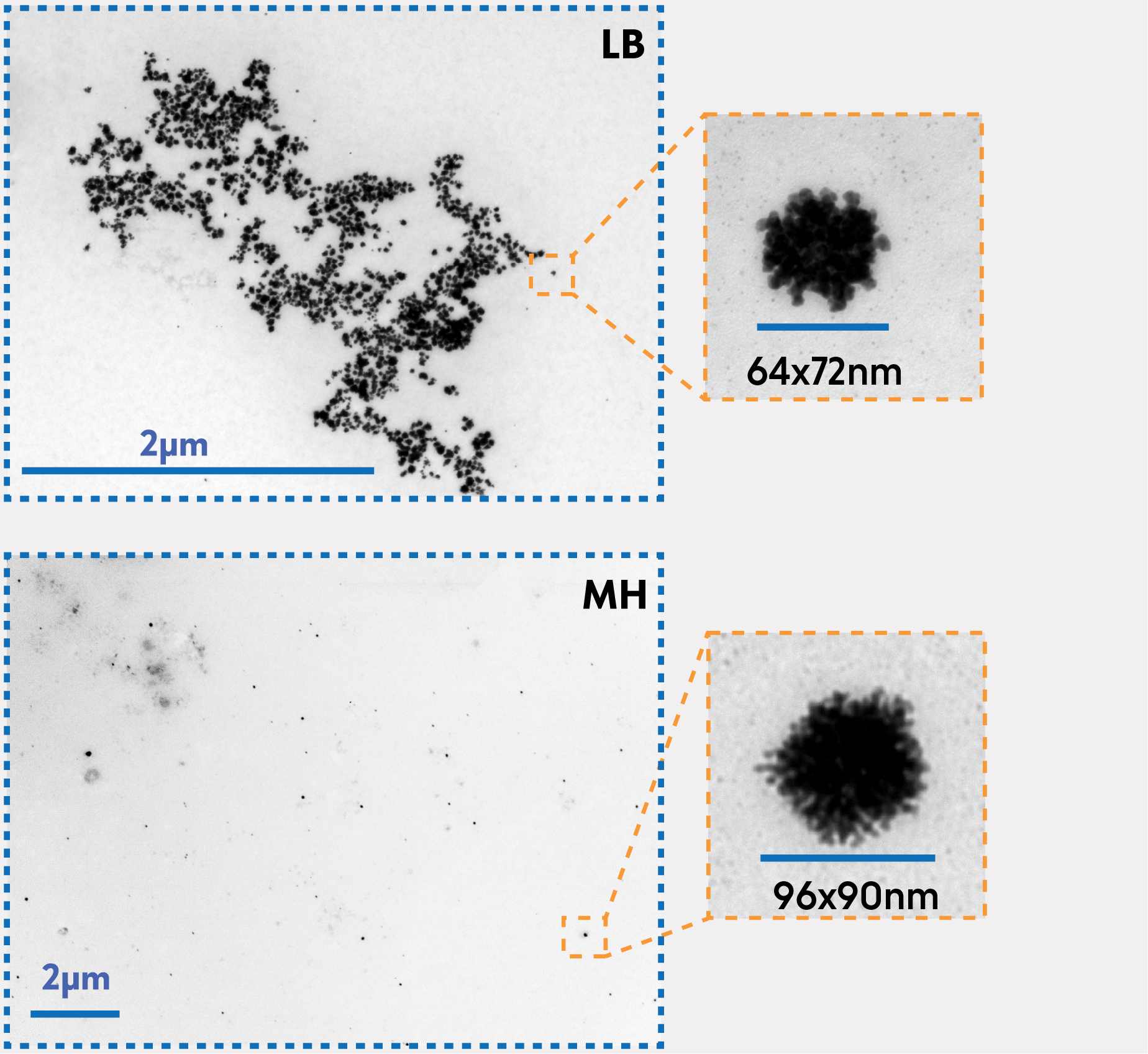
The optimal medium for nanoparticle production was chosen based on the best absorbance at 800 nm and further confirmed with Transmission Electron Microscopy (TEM). An overview of the main findings is presented below, see the bar plots with A800 nm of different media (Fig. 5) and TEM micrographs (Fig. 6). A complete description of our results can be found on the Results page.

Learn
MH supernatant without the lysate was found to have the best capacity for producing nanoparticles with the desired morphology (see Fig. 6 for spiky spheres) and a plasmon peak at 800 nm. Therefore, this engineering cycle helped us establish another parameter for Binanox: the optimal medium for bacterial growth of nanoparticle-producing E. coli.
It was also noticed that the control samples with media, silver, and gold salts could also produce nanoparticles. Therefore, to test which constituents of the media were responsible for the reducing capacity, another experiment was set up, as described on the Results page.
Cycle 3. Boost Binanox with synthetic biology

Motivation
Nanoparticle synthesis can be further improved by overexpression of proteins that contribute to bioreduction of metal salts.

Design
Adjust the nanoparticle synthesis protocol to span over 24h and test 4 common growth media.

Build & Test
Test the effect of adding a pellet with overexpressed proteins to the bioreducing mix on nanoparticle synthesis.

Learn
Adding the cell lysate to the bioreduction mix lessens its reducing capacity. Consider ensuring secretion of the overexpressed proteins.
After establishing the optimal medium and treatment of the biological system for nanoparticle production, we proceeded to the next step: further improving the biomanufacturing of nanoparticles by adding specific metal-reducing proteins. As seen in the literature, Cupriavidus metallidurans proteins CopA 9, 10 and NapA 11, 12, 13 have been studied in nanoparticle synthesis. In addition, the cup1 gene from Candida albicans was also shown to improve nanoparticle synthesis14. The beneficial effect of these proteins is mainly achieved through the increase in the reduction capacity of the mix. This typically leads to a higher yield and can influence the morphology4.
To test if the overexpression of these genes has a beneficial effect on nanoparticle synthesis in Binanox, we designed plasmids with the genes of C. albicans and C. metallidurans. Unfortunately, the transformations could not be performed within the limited lab time. You can read more about this on the Results page. This led us to substitute these genes with analogs from E. coli obtained from the ASKA library15. This way, we tested copA, napA, cueO and, additionally, melA, which was also found to have the potential to improve nanoparticle synthesis16.
ASKA Library: A complete Set of E. coli K-12 ORF Archive. This library contains E. coli K-12 clones with every individual gene of E. coli, fused with a His-tag and, optionally with GFP. The genes are also framed by an IPTG-inducible promoter and Sfil restriction sites. At Binanox, we utilized this library for overexpression of E. coli proteins CopA, NapA, CueO and MelA.
Design
To test the effect of the selected genes on nanoparticle production, we isolated the ASKA pCA24N plasmids from the library-standard E. coli K12 and cloned them into our E. coli BL21 strain. This allowed us to overexpress these genes. The proteins were all expressed intracellularly, so after IPTG induction, the pellet of a liquid culture would contain a high concentration of the protein of interest. After this, the pellet was resuspended in a smaller volume and lysed through sonication. Finally, this pellet lysate containing the overexpressed protein of interest was added to the MH supernatant of E. coli BL21 with an empty pET16b plasmid (introduced for antibiotic resistance). We then added silver and gold salts to this reaction mix to study nanoparticle formation.
The expected effect of all proteins on nanoparticle synthesis was a higher yield due to an increase in the reduction capacity of the mix. However, a change in the reaction mix can lead to changes in the synthesized nanoparticles' morphology, size distribution, and composition4. In this cycle, we directed our attention to the yield of nanoparticles with an absorbance at 800 nm, expressed in the intensity of A800 nm. Such an inference could be made after a supplementary experiment that found a direct correlation between nanoparticle concentration and absorbance (for more information, see the Results page).
Build & Test
Upon confirmation of the IPTG-induced overexpression of the proteins (see Results page for SDS gel), nanoparticle synthesis was performed according to the methodology described above. Additionally, when MelA was used for synthesis, L-DOPA was added, as MelA converts L-DOPA into the reducing agent melanin. Finally, absorption spectra were measured and compared to the controls without adding the proteins (Fig. 7).
A comparison was held between wildtype (control) and samples that contained the overexpressed proteins. Error bars represent standard error (n = 3). See Results page for full spectra, including those of control samples.
Learn
All the tested ASKA genes showed a significant decrease in A800 nm when added to MH (p < 0.05). By contrast, when added to WT E. coli BL21 supernatant, no significant difference (p > 0.20) was observed compared to the control. Moreover, most samples leaned towards a decrease in A800 nm, except for MelA with L-DOPA. MelA-containing lysate resulted in a significant increase for A800 nm compared to its control (p < 0.0005), which contained a lysate of WT. However, this increase was insignificant compared to the control without any lysate (p = 0.317). Observing that the absorbance becomes lower upon the addition of lysate is peculiar.
A potential clarification for this decline in nanoparticle producing capacity is the interaction of the overexpressed proteins with other compounds in the reducing mix, which causes the overall mix to be less effective in synthesizing nanoparticles. Such compounds could be other oxidizing agents that compete with metal ions. Alternatively, proteases could be disturbing the functionality of the reducing peptides. Both scenarios lead to a decrease in nanoparticle yield, as the reduction of metal ions experiences a significant drop. A more elaborate discussion of this prospect is discussed on the Results page. In addition, the integration of these results into Binanox is discussed in Cycle 4.
Cycle 4. Further improving the Binanox system with nitrate-containing media

Motivation
Nitrate-containing media induces the production of metal-reducing enzymes in E. coli.

Design
Test the influence of E. coli supernatant grown on MH-Nitrate on nanoparticle synthesis.

Build & Test
Compare the effect of supernatant of E. coli grown on MH vs MH-Nitrate on nanoparticle synthesis.

Learn
Nitrate in the medium does not bear a significant effect on nanoparticle synthesis when cell lysate is used.
The last step we undertook to optimize our biomanufacturing process was ensuring a better reducing power of the supernatant, to which we added the lysate with overexpressed proteins. According to a paper by Gurunathan et al.8, the presence of nitrate in the medium during growth increases the reducing capacity of the supernatant due to an increase in nitrate reductase production by the organism8. In addition, nitrate reductases, such as NapA, are responsible for silver nanoparticle synthesis17. Following this train of thought, we decided to test whether adding nitrate to MH would improve the yield of nanoparticle production by the supernatant-lysate mix.
Design
We compared the performance of the total ASKA-protein-containing bioreduction mix with the supernatant of MH to the supernatant of MH-Nitrate. We hoped to see an increase in the A800 nm of the obtained samples, which would signify that more nanoparticles with the desired properties are being synthesized.
Build & Test
400-1000 nm absorption spectra were measured for all reducing blends. The addition of the supernatant of MH was compared to the addition of the supernatant of MH-Nitrate. Fig. 8 shows the A800 nm for all samples that contained gold and silver salts.
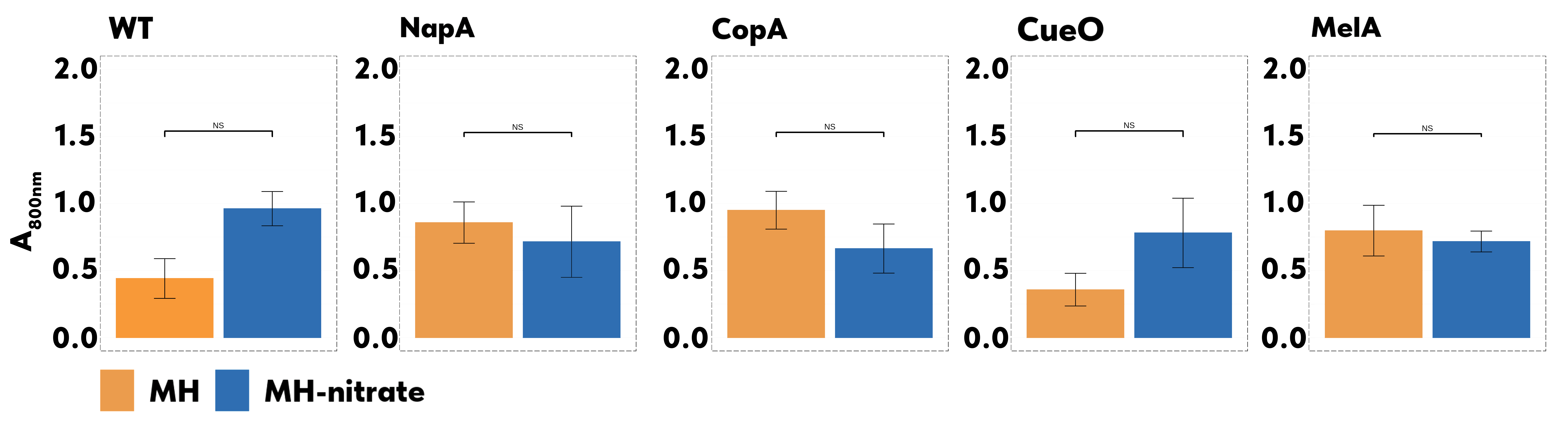
Learn
None of the samples showed a significant change in the absorbance at 800 nm between MH and MH-Nitrate supernatant (p > 0.05). This was an unexpected result, but we see potential explanations. One possibility is that nitrate's effect on these systems was insignificant due to the overriding reducing capacity of the overexpressed ASKA genes, which deems the influence of other reducing agents negligible. Another possibility is that similarly to the experiment in Cycle 3, the presence of the cell lysate in the mix introduced other oxidizing agents into the sample, which decreased the metal-reducing capacity of supernatant grown on MH.
The main conclusion from Cycles 3 and 4 is the necessity to eliminate the probable detrimental effects of the lysate on the reducing blend. Therefore, an alternative way to add the overexpressed proteins to the mix is necessary, for example, by tagging the proteins with secretion tags. This way, they would make up a part of the supernatant, which is expected to show an even better performance in reducing nanoparticles.
Conclusion
Going through the four engineering cycles, we got to learn more and more about the optimization of our nanoparticle biomanufacturing system Binanox. This invaluable information for process design could not have been obtained without getting negative results from the setup experiments, learning from them, and adjusting the system accordingly. Although each cycle helped us obtain important results individually, the first two cycles were used as input for the setup of the following two cycles. Moreover, all of the conducted experiments led us to conduct experiments to further investigate the matter of enhancing the synthesis of PTT-optimized nanoparticles.
Future experiments
Introduction
As an iGEM project only has a duration of a few months, there is not enough time to fully produce a complex product like a manufacturing process or a therapy. Hence, our team has looked further than just the end of iGEM and proposed future experiments. These future experiments are inspired by conversations with stakeholders and show what steps need to be taken to establish a final product. We have identified three areas of research that will require further study.
Biomanufacturing
Goal: Improve upon our current production process.
Design - Cell-free: During our lab time we did not use a protease inhibitor to prevent the degradation of our reducing proteins when lysing the bacterial cells. This likely caused the unfortunate results in our cell-free experiments. Therefore, we would like to do more experiments to further explore the cell-free production using a protease inhibitor.
Design - Simultaneous reduction: To improve upon our production design, we could in the future use ion-specific enzymes. With the current method of our production process and that of chemical synthesis, gold is reduced faster than silver. To enable simultaneous reduction of gold and silver the use of ion specific enzymes could be the key18.
Design - Upscaling: We should adapt our production process in order to obtain a consistent nanoparticle yield over different batches. This includes doing upscaling experiments, whereby we will aim to obtain stable concentrations in 10 L volumes. These results are then scaleable to 1000 L bioreactors.
Design - Isolation and Sterilization: Furthermore, we need to find an optimal isolation and sterilization method to obtain clean nanoparticles that can be utilized in medical applications. To obtain isolated nanoparticles we could use an ultracentrifuge or nanocentrifuge dependent on the weight of our nanoparticles. In regard to obtaining sterilized nanoparticles, potential methods are UV light and filter sterilization. Some of these methods have been tested. However, an expansion on these experiments would be vital to the success of our nanoparticles.
Analysis
Goal: Further characterize nanoparticles.
Design: Nanoparticle behavior is highly dependent on their size, shape and surface area19. Therefore, there are some additional measurements that we could do to get a better understanding of our nanoparticles. As can be read in Human Practices, we spoke to an expert about the NTA, DLS and Nanocet-1. Each of these measures can determine nanoparticle size. Furthermore, if we decide to explore the opportunity of a diagnostic application further, we should determine the density of our nanoparticles. We could use a method like Small Angle X-ray Scattering (SAXS)20. Lastly, literature indicated the collapsing of spikes due to heat. It is interesting to discover at which temperature and duration of heating this will occur and whether it will influence our implementation.
Toxicity
Goal: To accurately model in vivo responses to our nanoparticles.
Design: To make sure our nanoparticles are safe for in vivo
use, we need to determine whether coating has a significant effect on
toxicity. To further elucidate toxicity and photothermal therapy effectiveness
in vitro experiments using both cancer and non-cancer cell lines are
desired. We will have to test different concentrations of nanoparticles and the
duration of heating during the in vitro experiments. Additionally, to
comply with EMA regulations we will have to investigate cyto- and genotoxicity.
During in vivo experiments, we will also have to investigate the
ability of our nanoparticles to infiltrate tumor tissue and the effects of
antibody-nanoparticle conjugation on surface plasmon resonance and the
effectiveness of PTT.
- Sudha, P. N., Sangeetha, K., Vijayalakshmi, K., & Barhoum, A. Nanomaterials history, classification, unique properties, production and market. Emerging Applications of Nanoparticles and Architecture Nanostructures, 341–384 (2018)
- Cheng, L. C., et al. Seedless, silver-induced synthesis of star-shaped gold/silver bimetallic nanoparticles as high efficiency photothermal therapy reagent. J. Mater. Chem. 22, 2244–2253 (2012).
- Yeh, Y. C., Creran, B., & Rotello, V. M. Gold nanoparticles: preparation, properties, and applications in bionanotechnology. Nanoscale 4, 1871–1880 (2012).
- Rodrigues, T. S. et al. Synthesis of Colloidal Metal Nanocrystals: A Comprehensive Review on the Reductants. Chemistry – a European Journal, 24, 16944–16963 (2018).
- Zhang, D. Green Synthesis of Metallic Nanoparticles and Their Potential Applications to Treat Cancer. Frontiers 8 (2020).
- Davey, K. Modelling the combined effect of temperature and pH on the rate coefficient for bacterial growth. International Journal of Food Microbiology 23, 295–303 (1994).
- Shahverdi, A. R., Minaeian, S., Shahverdi, H. R., Jamalifar, H., & Nohi, A. A. Rapid synthesis of silver nanoparticles using culture supernatants of Enterobacteria: A novel biological approach. Process Biochemistry 42, 919–923 (2007).
- Gurunathan, S. et al. Biosynthesis, purification and characterization of silver nanoparticles using Escherichia coli. Colloids and Surfaces B: Biointerfaces 74, 328–335 (2009).
- Bütof, L. et al. Synergistic gold–copper detoxification at the core of gold biomineralisation in Cupriavidus metallidurans. Metallomics, 10, 278–286 (2018).
- Reith, F. et al. Mechanisms of gold biomineralization in the bacterium Cupriavidus metallidurans. Proceedings of the National Academy of Sciences 106, 17757–17762 (2009).
- Vaidyanathan, R. et al. Enhanced silver nanoparticle synthesis by optimization of nitrate reductase activity. Colloids and Surfaces B: Biointerfaces 75, 335–341 (2010).
- Khodashenas, B. Nitrate reductase enzyme in Escherichia coli and its relationship with the synthesis of silver nanoparticles. Journal of Research in Science, Engineering and Technology 3, 26–32 (2019).
- Fu, J. K., Zhang, W. D., Liu, Z. Y., Yao, B. X., & Weng, S. Z. Characterization of adsorption and reduction of noble metal ions by bacteria. Chin J Chem Univ 20, 1452–1454 (1999).
- Yuan, Q. Enhanced Silver Nanoparticle Synthesis by Escherichia Coli Transformed with Candida Albicans Metallothionein Gene. Material 12, 4180 (2019).
- Kitagawa, M. et al. Complete set of ORF clones of Escherichia coli ASKA library (A Complete Set of E. coli K-12 ORF Archive): Unique Resources for Biological Research. DNA Research 12, 291–299 (2006).
- Tsai, Y. J. et al. Biosynthesis and display of diverse metal nanoparticles by recombinant Escherichia coli. RSC Adv. 4, 58717–58719 (2014).
- Anil Kumar, S. et al. Nitrate reductase-mediated synthesis of silver nanoparticles from AgNO3. Biotechnology Letters 29, 439–445 (2007).
- Esmail, R., Afshar, A., Morteza, M., Abolfazl, A. & Akhondi, E. Synthesis of silver nanoparticles with high efficiency and stability by culture supernatant of Bacillus ROM6 isolated from Zarshouran gold mine and evaluating its antibacterial effects. BMC Microbiol 22, (2022).
- Jeevanandam, J., Barhoum, A., Chan, Y. S., Dufresne, A. & Danquah, M. K. Review on nanoparticles and nanostructured materials: history, sources, toxicity and regulations. Beilstein journal of nanotechnology 9, 1050–1074 (2018).
- Minelli, C. et al. Measuring the size and density of nanoparticles by centrifugal sedimentation and flotation. Analytical methods 10, 1725–1732 (2018).

