The Problem
Invasive Fungal Infections

Nosocomial infections pose a major threat to immunocompromised and critically-ill patients as well as hospital staff (Kariyawasam et al. 2022). Candidiasis is an invasive fungal infection caused by the opportunistic pathogen Candida albicans that often presents in hospitals.
Antimicrobial Resistance
Antimicrobial resistance remains a major global threat and hospitalizations by COVID-19 are a contributor to the spread of multidrug-resistant pathogens throughout hospitals (Xia 2022). COVID-19 patients have a significant risk of developing antibiotic/antifungal resistant co-infections, including invasive fungal infections.
Claymore
Claymore is an engineered organism designed to detect and dispatch pathogenic organisms in vivo. Our project this year is a proof of concept with the test organism Candida albicans. Claymore has two major advantages over traditional antibiotic/antifungal use:
1. Highly Targeted

Claymore is outfitted with a detection system so antibiotics/antifungals will only be produced in the presence of the pathogen. A highly targeted system will prevent antibiotic/antifungal production in areas where the pathogen is not present which reduces damage to the microbiome and prevents the development of antimicrobial resistance.
2. Self-Sufficient

Because Claymore is a living organism, it is capable of reproducing at the site of infection. As a result, only a single dose is necessary to thwart an infection rather than regularly scheduled doses. In addition, Claymore will be active until the target pathogen is killed, reducing the risk of antimicrobial resistance developing after treatment or if a treatment is incomplete.
Read more about Claymore on the project description page.
Our Project and Parts Design
Our project is broken down into two parts, a detection system to sense the pathogen and elimination system that produces a toxin to kill the target pathogen. For this year, we are using the HPA system as a detection system and the bacillomycin D biosynthetic pathway as an elimination system.
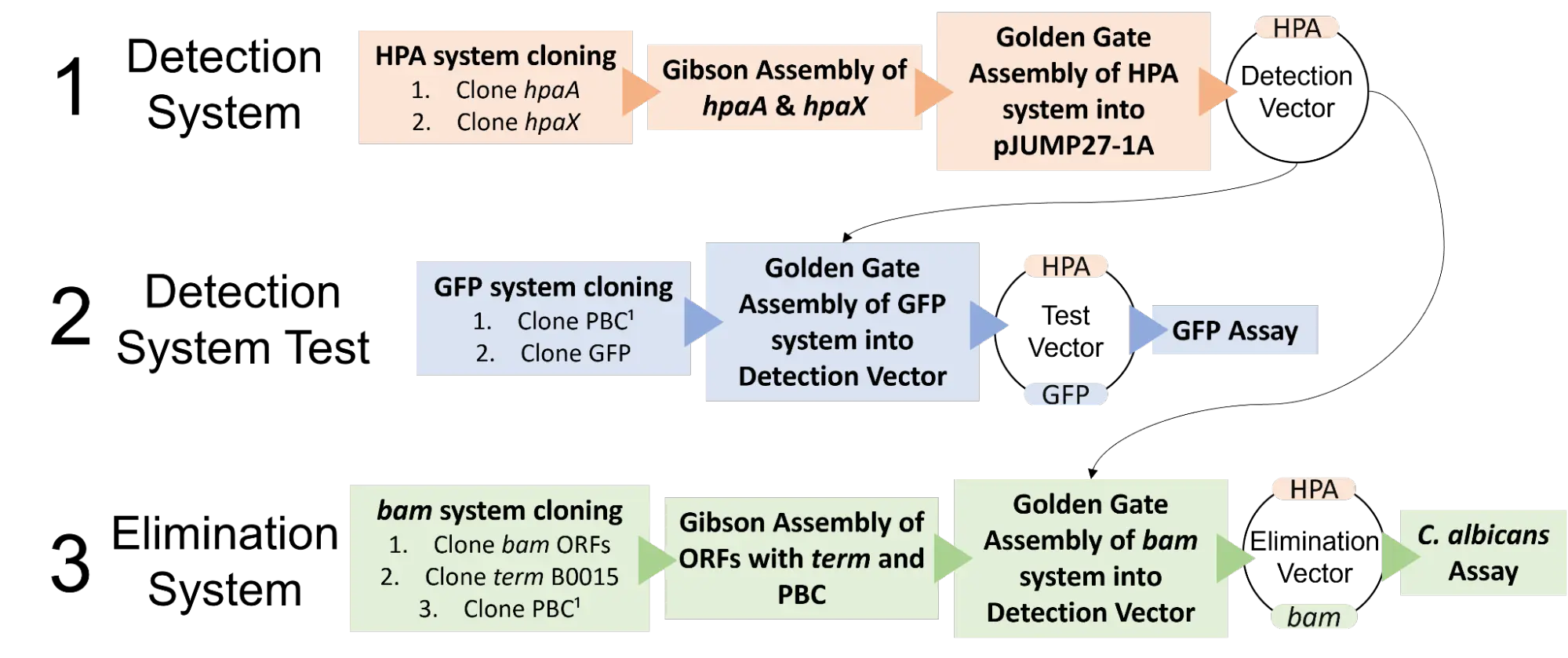
¹PBC is a promoter controlled by the HPA system.
Our project design page and parts page have more in-depth descriptions of our project altogether, please check them out!
The Future of Claymore
While we adapted Claymore this year to target and eliminate C. albicans, we hope to develop a system that can be adapted to target various pathogenic fungi. We plan to maintain the HPA system while incorporating varying detection modules and elimination systems for different pathogens. We also hope to optimize Claymore to include components such as a kill switch and synergistic antimicrobial agents.
Sponsors
The University of Alberta Faculty of Science donated $1000 to our team this year, allowing our lab to run smoothly for the duration of our research period.
Fisher Scientific provided our team with vital pieces of equipment that were nearly impossible to source on campus. Their equipment loan allowed us to move forward in the project at a critical time. The equipment loan consisted of the following items: Benchtop mini centrifuge, Mini AGE system, Lab timer, Mini vortex mixer, Benchtop tube roller, Benchtop microplate shaker, Serological pipette manual pipette aid, rechargeable accuject pipette aid, P20, 200, 1000 (x2) air displacement pipettes.
SnapGene provided free access to their premium software for the duration of our research period which was beyond critical in accomplishing the dry and wet lab portions of the project.
Citations
Kariyawasam RM, Julien DA, Jelinski DC, Larose SL, May ER, Conly JM, Dingle TC, Chen JZ, Tyrrell GJ, Ronksley PE, et al. 2022. Antimicrobial resistance (AMR) in COVID-19 patients: A systematic review and meta-analysis (November 2019-June 2021). Antimicrob Resist Infect Control. 11(1):45. doi 10.1186/s13756-022-01085-z
Xia J, Wang Z, Li T, Lu F, Sheng D, Huang W. 2022. Immunosuppressed patients with clinically diagnosed invasive fungal infections: The fungal species distribution, antifungal sensitivity and associated risk factors in a tertiary hospital of Anhui Province. Infect Drug Resist. 15(1):321-333. doi 10.2147/IDR.S351260.



